Thrust Area (TA) Projects
Thrust Area 1 Thrust Area 2 Thrust Area 3
Thrust Area 1
PhenoProfiling

Science Objectives:
- Functionally enrich microbial communities and generate multi-omics to correlate biochemical mechanisms to activity.
- Integrate PhenoProfiling with Thrust Areas 2 and 3 to develop models for phenotype prediction and interspecies interactions.
- Evaluate enriched consortia for biochemical and molecular shifts as evidence of phenotypic changes related to soil carbon sequestration.
Profiling the Structural Proteome to Define Phenotype

Science Objective:
- Apply structural proteomics technologies to map the molecular interactome.
Multi-PTM Profiling for Molecular Phenotyping

Tong Zhang
Science Objective:
Nanobodies to Control Phenome Transient Forms

Amy Sims & Samantha Powell
Science Objective:
- Use stimuli-specific, synthetic nanobodies to target functional mediators without prior knowledge of the response networks or manipulating the biological system.
Thrust Area 2
Microbial Community Engineering to Reduce Carbon and Nitrogen Footprints in Biomanufacturing
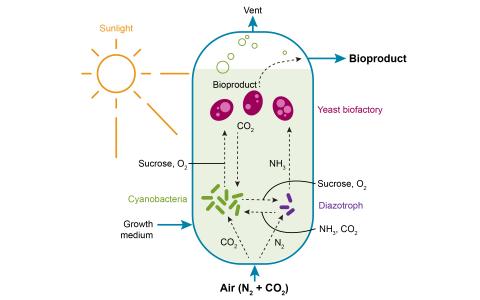
Pavlo Bohutskyi & Kyle Pomraning
Science Objective:
- Establish model synthetic microbial communities to understand the rules regulating their biological function in order to utilize them as next generation bioproduction platforms capable of reducing carbon and nitrogen footprints in biomanufacturing processes.
Controlling Host Cell Functions to Block Virus Replication
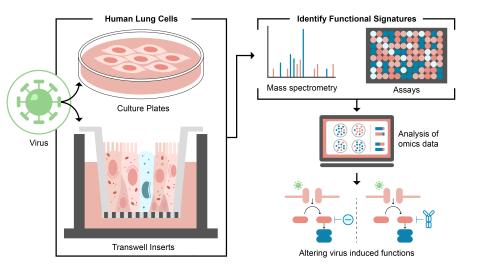
Science Objective:
- Identify and control host functions hijacked during viral infection through use of PNNL ‘omics technologies and modeling capabilities.
CarbStor
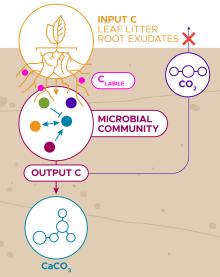
Science Objective:
- Build and understand model communities that show carbon storage phenotypes
Developing Genetic Code Expansion in Diverse Bacteria for Carbon Sequestration Applications

Elise Van Fossen & Joshua Elmore
Science Objectives:
- Incorporate genetic code expansion machinery into diverse bacterial species and engineer bacteria to secrete cross-linkable peptide precursors of synthetic peptidyl lignin
- Develop tools for genetically manipulating other bacteria in the Predictive Phenomics Initiative
- Generate chemically customized recombinant proteins for phenotype profiling
Thrust Area 3
PTM-Psi: A Python Package to Facilitate the Investigation of Post-Translational Modification on Protein Structures and their Impacts
Science Objective:
- Develop new theory and tools that leverage evolutionary perspectives and knowledge of the energetics of reactions to predict the most likely regulation in a given environment. These methods will accelerate exploration, modeling and understanding of cell regulation phenotypes.
Predicting Cell Regulation Phenotypes with Small Molecule Data
Ethan King
Science Objective:
- This project will develop computational methods to predict cell regulation phenotypes using small molecule and proteome data to understand outcomes in complex biological systems.
A Biologically Informed Bayesian Prediction Framework
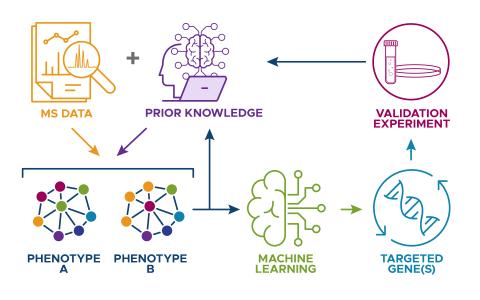
Science Objective:
- Develop a biologically informed ML model that integrates datasets from different studies, and leverages current biological knowledge in an automated manner, to improve predictions in biological data analysis.
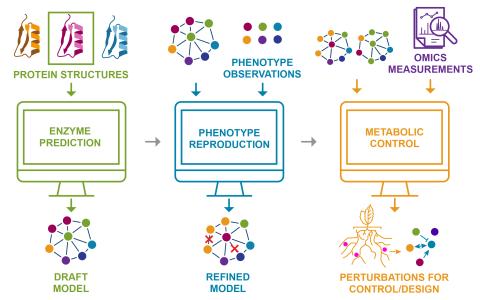
Model-Based Prediction and Control of Phenotypic Traits in Microbial Consortia
Jeremy Zucker & Vlad Petyuk
Science Objective:
- By developing explainable, predictive metabolic models of individual microbes, we aim to design consortia that convert light and abundant atmospheric gases into high-value molecules through microbial division of labor.
Predictive Phenomics Data Infrastructure
Grant Fujimoto

Developing Genetic Code Expansion in Diverse Bacteria for Carbon Sequestration Applications

Science Objective:
- We are constructing a streamlined approach to identify phenotype-relevant signatures by integrating various proteomics data. Leveraging protein structures and interaction networks, we will map structural changes and post-translational modifications to identify molecular drivers and subsequently emerged mechanisms for phenotypes. This method narrows down target sites, simplifying the search for pivotal points like drug or engineering targets. It offers efficiency, time savings, and it enhances scientific discovery over traditional screening methods.
Lab-Level Communications Priority Topics
Biology