Biological
Sciences
Division
Biological
Sciences
Division
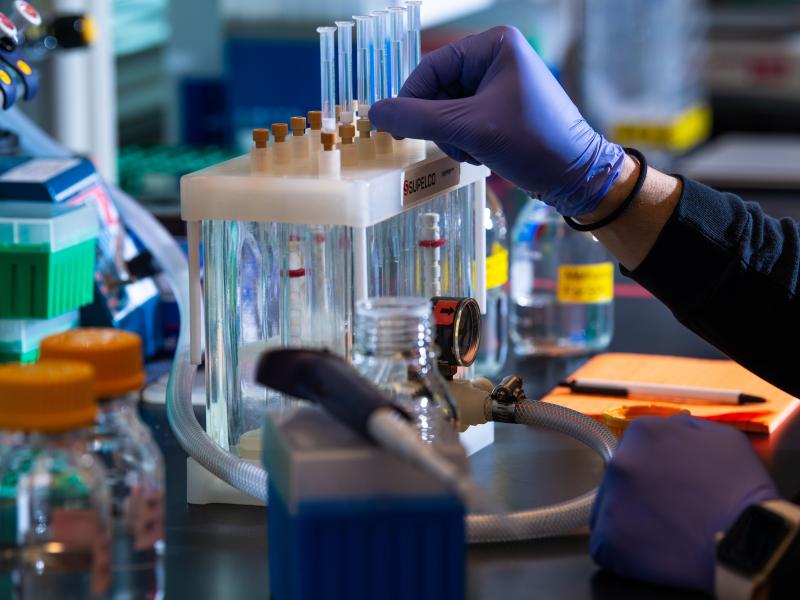
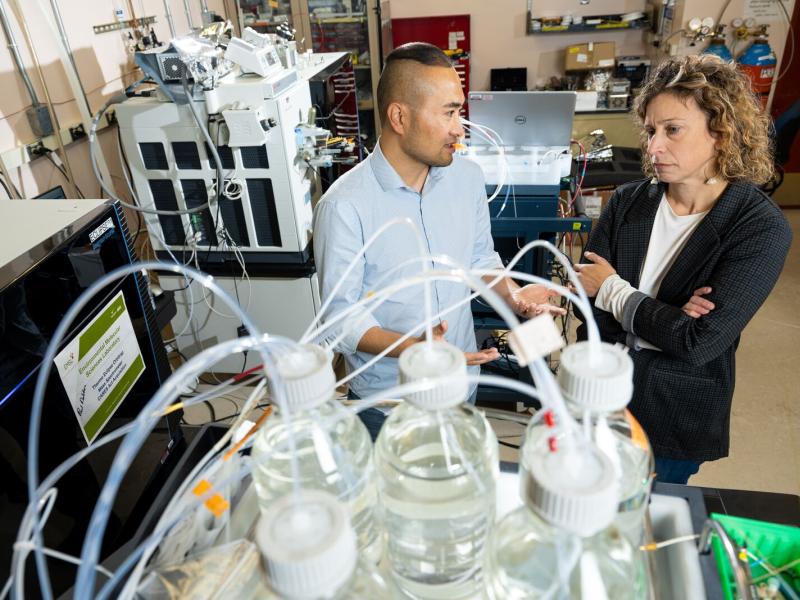
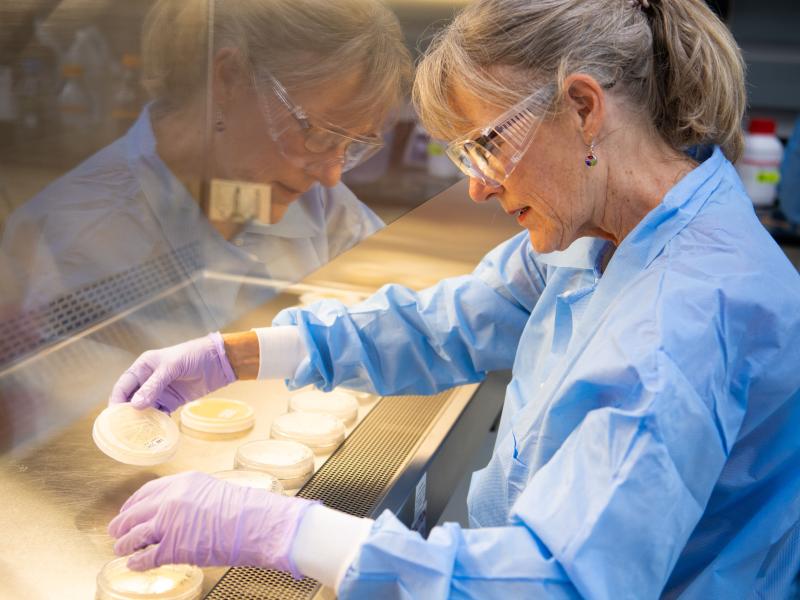
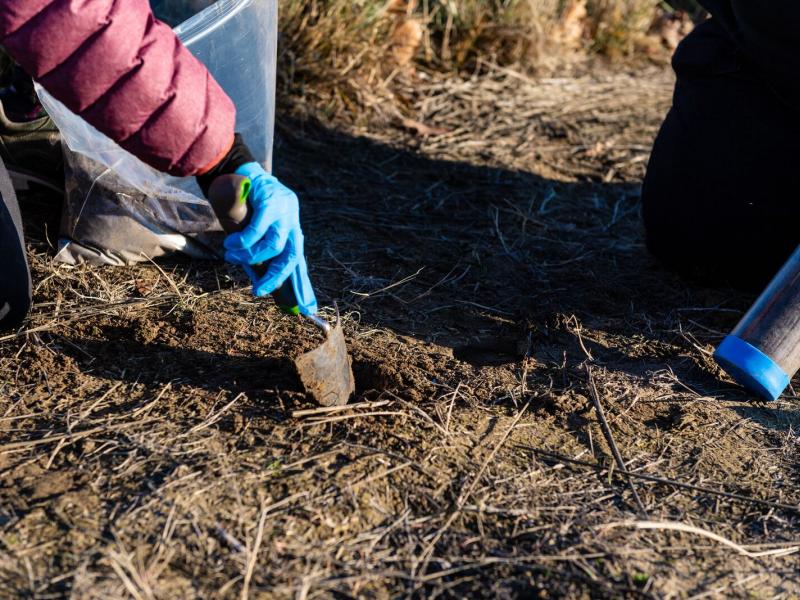
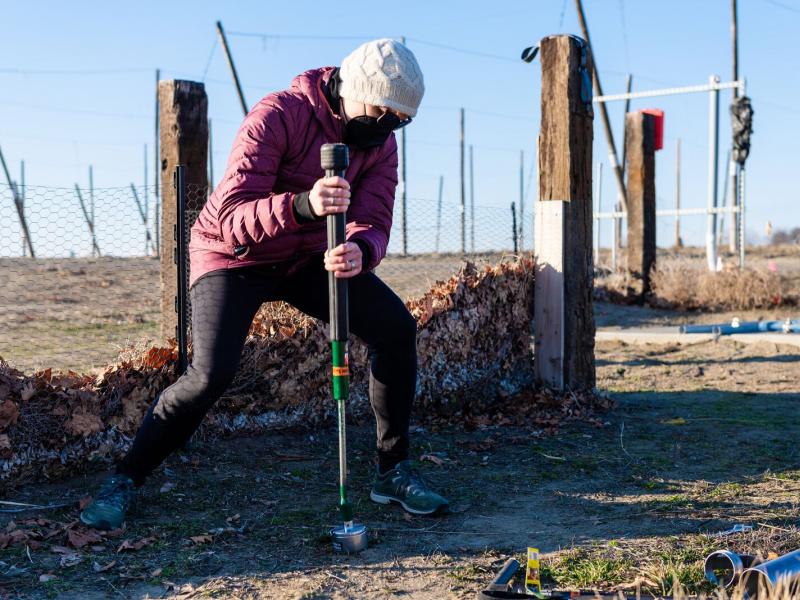
The Biological Sciences Division has 16 collaborative, interdisciplinary teams working to understand how plants, microbes, and algae function in a way that helps predict their response to environmental change and their potential for energy production. Insights into the workings of human tissues reveal how toxins and pathogens could influence human health. Our research inspires biology-based advances to tackle some of the biggest challenges our world faces in ecosystem sustainability and human health. Learn more about how PNNL teams explore these areas using instrumentation that provides large-scale, molecular-level information and computational tools to efficiently process vast amounts of data.
Biological Systems Science
Computational Biology
Integrative Omics
Featured News & Highlights
The Biological Sciences Division has 16 collaborative, interdisciplinary teams that inspire biology-based advances to tackle some of the biggest challenges our world faces in ecosystem sustainability, bioenergy, human health and national security.